Hi, and welcome to this video on Punnett squares!
Some of the terminology you’ll come across when dealing with genetics can be confusing, so we’ll start by defining some terms and then put them all together to see how they relate to each other. Let’s get started!
Reviewing Terminology
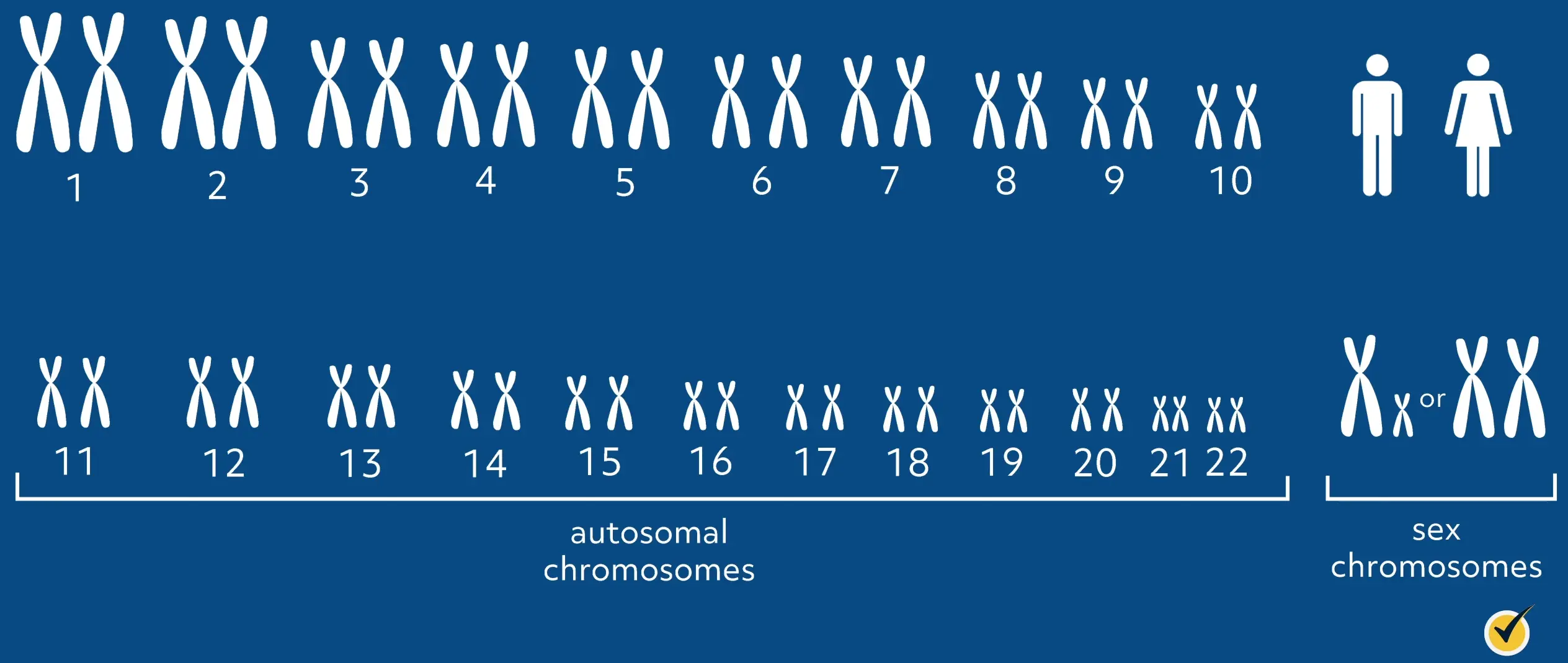
First, let’s remember that human somatic cells have 22 pairs of autosomal chromosomes–one from each parent–and one pair of sex chromosomes to total 23 pairs of chromosomes for a wildtype individual. This gives humans a total of 46 chromosomes. Chromosomes are made up of DNA that is tightly coiled around proteins that is called histones.
Genes are pieces of DNA found on chromosomes that can act as codes for a specific protein. Our somatic cells are diploid, meaning we have two copies of each gene, one from each parent. These genes influence all kinds of traits, but how do we know which traits offspring will inherit? It depends on the random separation of alleles during meiosis and the random re-combination of these alleles during fertilization.
Alleles
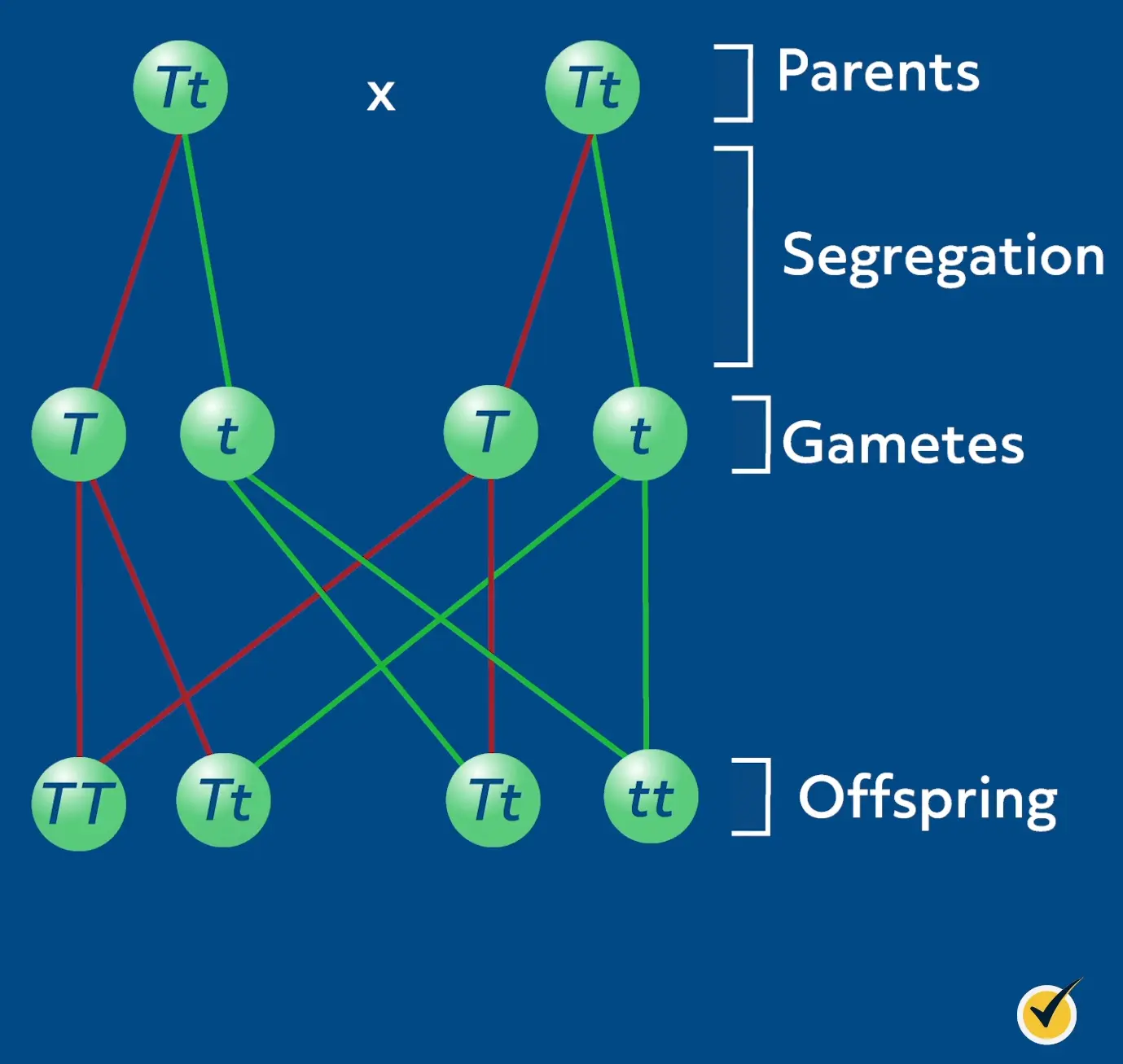
Alleles are sequence variants of a gene that produce a certain observable trait, or phenotype. We have one allele from mom and one allele from dad for each trait. Mendel’s law of segregation states that the parents’ allele pairs separate randomly and only one allele from each parent is passed to the offspring. This particular cell in the offspring has two alleles again, and it’s the combination of these alleles, or genotype, that determines the observable trait (phenotype).
Let’s take height for an example. In reality, there are many factors that contribute to height differences in humans, but for the sake of this example let’s say a person is either tall or short. Remember that all somatic cells contain two alleles for each trait, one on each chromosome. So the gene for height is going to be located in the same spot on each chromosome from your mom and dad. Even though this gene is located in the same spot and codes for the same height trait, the actual sequence of this part of the gene on each chromosome can vary.
Remember, this varied sequence is an allele. So in our height gene example, the gene that codes for height on one chromosome may have a sequence that reads like this: ACGTC. Let’s say this allele codes for “tall.” The gene that codes for height on the other chromosome has a sequence that reads: AGGTC. This allele codes for “short.” We can assign arbitrary upper and lowercase letters to our alleles to keep them organized. Let’s keep it simple and go with the letter “T” like this:
| Allele | Sequence | Phenotype |
|---|---|---|
| T | ACGTC | tall |
| t | AGGTC | short |
The possible allele combinations for this example are TT, Tt, or tt. These are known as the genotype, or the actual alleles that are present in the gamete for a single trait. Individuals that carry the same alleles are homozygous for that trait. So for height, you can be homozygous if you have alleles that read TT or tt. Individuals that carry different alleles are heterozygous for that trait, so they have one of each allele and have the genotype Tt.
Remember that a phenotype is the physical expression of your genes, or how the trait is observed, and we derive the phenotype from the genotype. In our example we labeled the phenotype “tall” with the capital letter T. Any time an allele is labeled with a capital letter it means it’s the dominant allele. A dominant allele will win over a lowercase, or recessive allele, in modes of inheritance.
So let’s say we have two parents and we want to know what the outcome of their offspring will be. We know that in meiosis, each parent gives one allele copy at random to each haploid gamete it forms. When this gamete fuses with a gamete from the other parent, it forms a zygote.
Using Punnett Squares
To predict the probability of an offspring having a particular genotype and phenotype from two parents, a Punnett square is a really good tool to use. So let’s make a Punnett square for our height trait. Since we’re only looking at one trait, this is called a monohybrid cross. Let’s say that both parents are heterozygous with the genotype Tt. Both parents would be tall because they each have the dominant T for tall allele that overshadows the recessive allele for short. When we want to trace the inheritance for a single trait to offspring we can use a simple 2×2 grid like this:
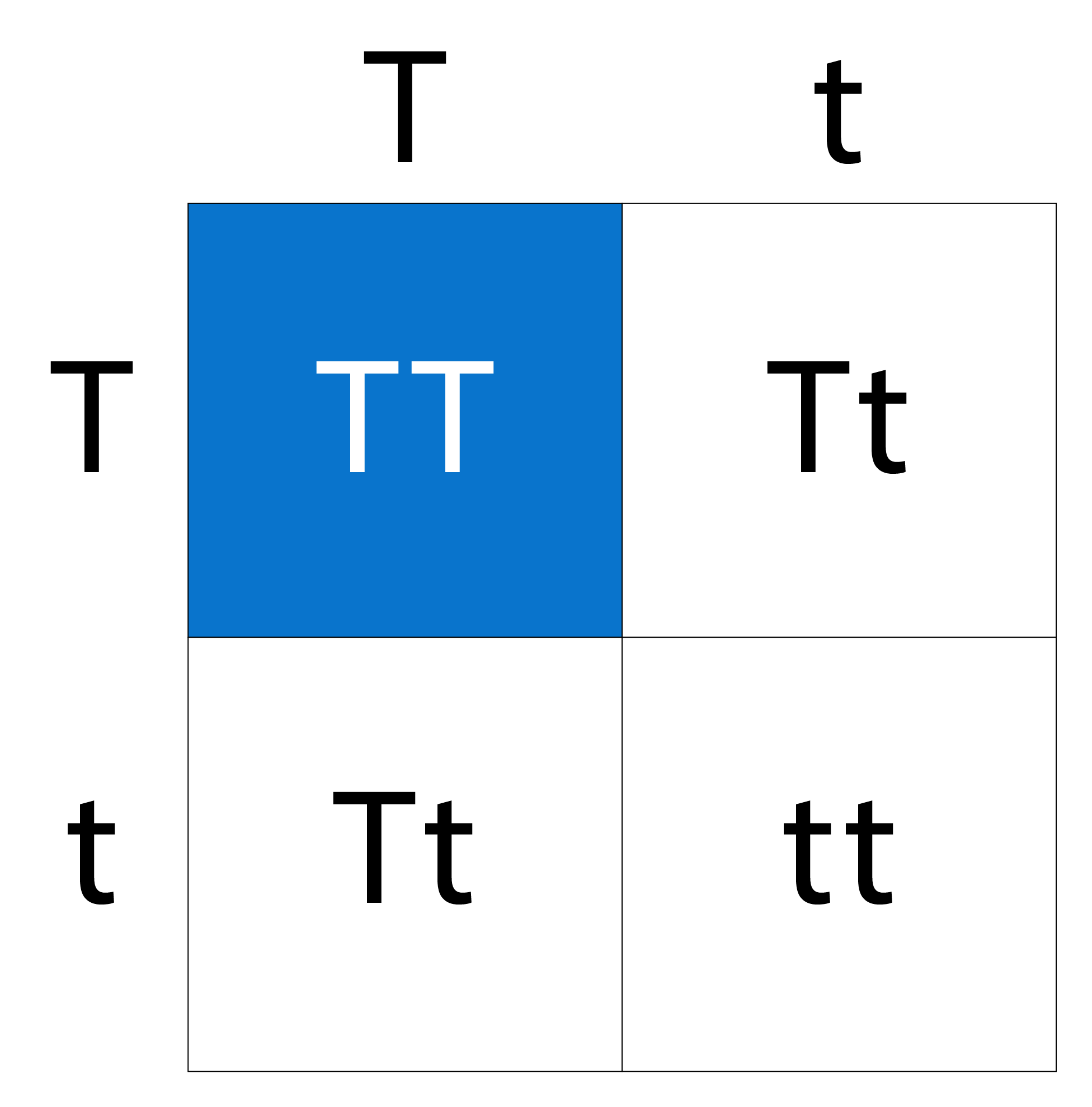
We’ll put Parent 1 across the top and Parent 2 along the side like so. Starting in the upper left corner, we’ll draw the alleles down and across the grid to fill each square. Let’s look at the results. These are the four possibilities of what the genotype of Parent 1 and 2’s offspring could look like. Since the tall (T) allele is the dominant allele, in the first quadrant there is a 25% chance the offspring will be homozygous dominant, or TT. This would mean the offspring’s phenotype is tall like the parents.
Two of the four quadrants possess the heterozygous dominant genotype, Tt, which means there’s a 50% chance the offspring would possess this genotype. Because the dominant allele, or T, is still in the pair, this means the offspring would still show a tall phenotype. Notice how the bottom right quadrant has two t’s. This genotype is known as homozygous recessive, because it contains two of the same recessive alleles (tt). Because one of the four quadrants contains this genotype, there is a 25% chance that the offspring will be short.
Patterns of Inheritance
Punnett squares like this also help us see certain patterns of inheritance. In our example, we had a 1:2:1 genotypic ratio for homozygous dominant, heterozygous dominant, and homozygous recessive, respectively. This ratio stays the same no matter how many offspring the parents decide to have. We can also see a phenotypic ratio from our cross that shows their offspring have a 3:1 chance of being tall or a 1:3 chance of being short. This ratio also stays the same no matter the number of offspring because the results represent percentages. We could say that 75% of these parents’ offspring will be tall or 25% will be short.
Dihybrid Cross
We can also use Punnett squares if we want to follow the inheritance pattern of two traits to their offspring. This is called a dihybrid cross–“di” meaning two. The important thing with dihybrid crosses is that they show that the inheritance of one trait doesn’t influence how another trait is inherited, meaning if I inherit the trait for being tall, it doesn’t necessarily improve or lessen my chances for inheriting a different trait. This is called Mendel’s law of independent assortment and we can test it with a 4×4 Punnett square.
Let’s use the same height trait from our monohybrid cross but this time let’s also consider the trait for having a widow’s peak where the P allele means there’s a widow’s peak and the p allele means there isn’t. Both parents will be heterozygous for both traits meaning they both have the genotype TtPp. We’ll set it up just like before by placing the possible allele pairings from each parent along the top and side, but this time we’ll do it for both traits. If you need help finding all the possible gametes, use the FOIL method where you take the first, outside, inside, and last letters for each trait and write them down. So our first parent would contribute TP, then tP, Tp, and tp, and Parent 2 could contribute the same.
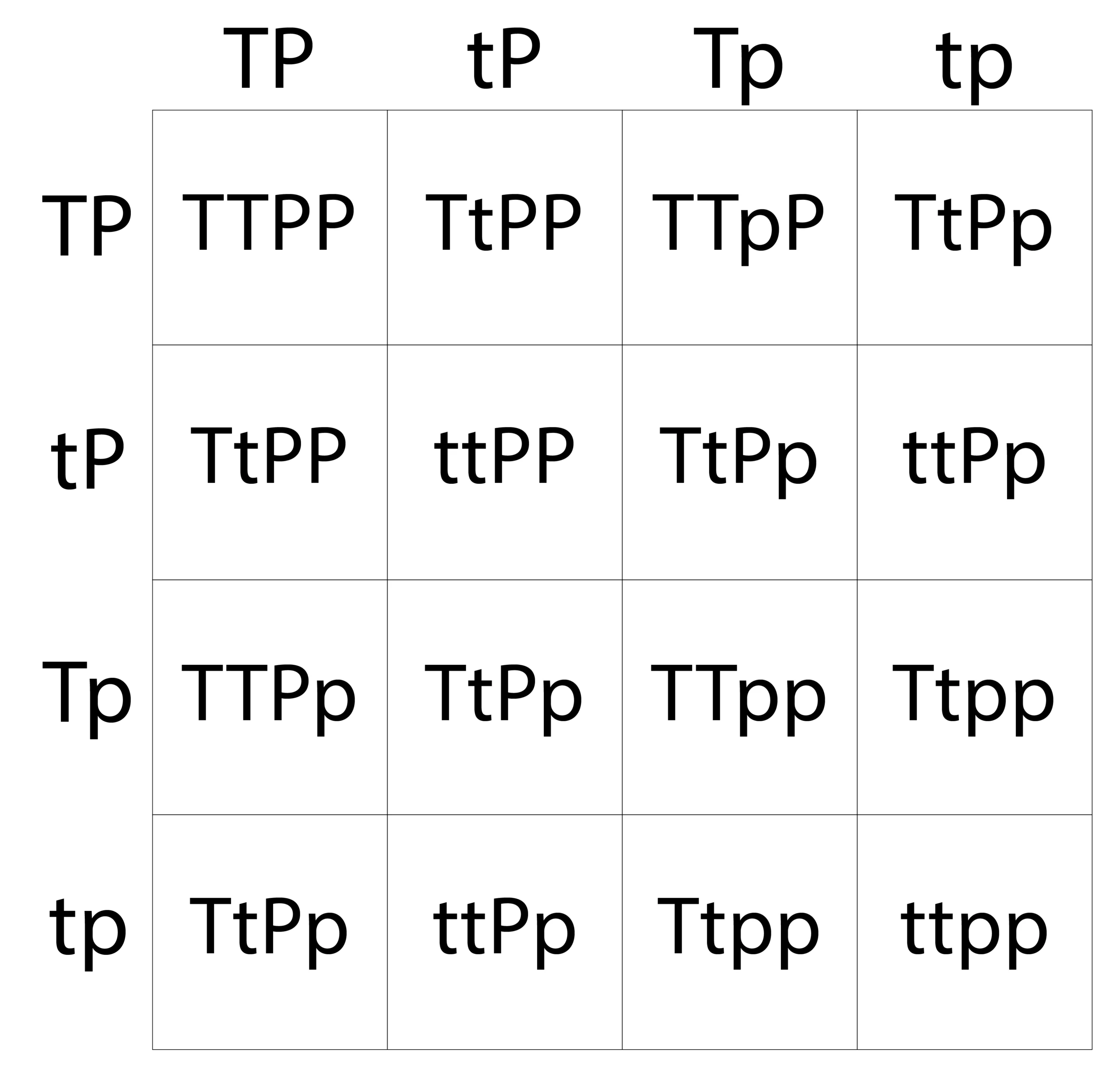
Now, let’s fill in all of the squares. When we bring the alleles down and across for each square, it’s helpful to keep the traits together when we fill in the squares. Now we can see a pretty clear pattern. We have nine offspring that will be tall and have a widow’s peak, three that are short with a widow’s peak, three that are tall and don’t have a widow’s peak, and one that is short with no widow’s peak. This is the classic 9:3:3:1 phenotypic ratio that we expect from a dihybrid cross. You can use the same method to do a testcross with different genotypes for the parents and their gametes if you want to see the different resulting phenotypes and ratios.
I hope this review was helpful! Thanks for watching, and happy studying!
